MLZ is a cooperation between:
 > Technische Universität München
> Technische Universität München > Helmholtz-Zentrum Hereon
> Helmholtz-Zentrum Hereon
 > Forschungszentrum Jülich
> Forschungszentrum Jülich
MLZ is a member of:
 > LENS
> LENS > ERF-AISBL
> ERF-AISBL
MLZ on social media:

MLZ (eng)
Lichtenbergstr.1
85748 Garching
06.02.2024
Examining paramedic proteins
Proteins unfold when exposed to heat and thus not only lose their function, but can clump together and cause damage in organisms. Heat shock proteins (Hsp) bind to proteins before they unfold completely. A group from Japan has studied Hsp72 with neutrons for the first time and gained important insights into the course of these reactions.

Dr. Andreas Ostermann, instrument scientist, adjusts the bio-diffractometer at the Heinz Maier-Leibnitz Zentrum. © Andreas Heddergott, TUM
Anyone can observe what happens when egg white is heated while preparing eggs on the stove: When exposed to heat during frying or boiling, the proteins unfold and clump together: the egg becomes solid. This should not happen in living cells with the first sunshine in summer. With so-called heat-shock proteins, nature has created a remedy for this. Professor Takeshi Yokoyama’s group at the University of Toyama, Japan, wants to better understand the function of Hsp72, a special heat-shock protein that is also used in human cells, and has made the first measurement of this protein with neutrons at the BIODIFF instrument of the MLZ.
First aid through capture
Heat-shock proteins such as Hsp72 step in and provide first aid by capturing the unfolded proteins. As soon as the environmental conditions return to normal, the protein is released and can fold back into its original state. This cycle is possible because of the two domain structure of Hsp72. One domain, the SBD (substrate-binding domain) binds the unfolding protein (also known as the substrate). The other, the NBD (nucleotide-binding domain) binds the nucleotide ATP (adenosine triphosphate), which provides the energy for this cycle by being split into two smaller molecules: Into the ADP and a monophosphate group.
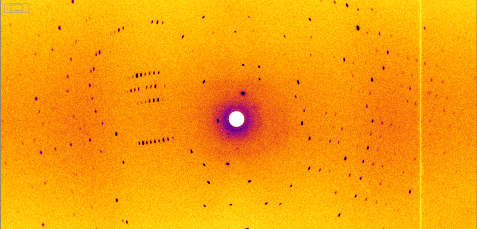

This is the result of the measurement on the BIODIFF instrument: A diffraction pattern of the heat shock protein. The researchers determined the three-dimensional structure of the heat shock protein and the water clusters from 378 such images. © Dr. Andreas Ostermann, FRM II / TUM
Neutrons show the positions of hydrogen atoms
For their measurements, Professor Yokoyama’s group examined the NBD part of Hsp72, to which an ADP is bound. The aim of the investigation with neutrons was to determine whether or not hydrogen is bound to the phosphate groups of the ADP and to the three sites that are critical for the cleavage. It turned out that this is not the case – an important piece of information for estimating the electrostatic conditions in the active center.
In addition, the team was able to identify three clusters of ordered water molecules in the structure. Neutrons are ideal for such investigations. “The orientation of water molecules in or near the active center can often only be determined using neutron scattering. “, says Dr. Andreas Ostermann, instrument scientist at BIODIFF.
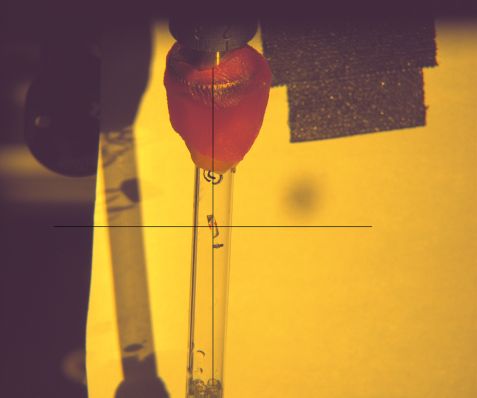

In the crosshairs, the protein crystal of the heat shock protein under investigation. If the proteins are obtained as crystals, scientists can use neutron scattering methods to unravel their structure. © Dr. Andreas Ostermann, FRM II / TUM
Findings help in the development of cancer drugs
“Using the measurements, we discovered that the stability of a certain water cluster at the active center, where the ATP is cleaved, strongly influences this process. The more disordered the water molecules, the higher the activity of ATP cleavage,” says Professor Yokoyama. The interplay between the activity and ordered water structures in the active center is very interesting. With a better understanding of this interplay, one can think about how to control the activity.
Malfunction of Hsp is related with diseases such as cancer. “Therefore, Hsp72 is also being recognized as a target protein for anticancer agents. The information about hydrogen atoms in Hsp72 obtained through the measurements at BIODIFF could contribute to the design and development of new anticancer drugs”, explains Professor Yokoyama.
Original Publication:
T. Yokoyama, S. Fujii, A. Ostermann, T. E. Schrader, Y. Nabeshima, and M. Mizuguchi
Neutron crystallographic analysis of the nucleotide-binding domain of Hsp72 in complex with ADP
IUCrJ (2022). 9, 562-572
DOI: 10.1107/S2052252522006297
Related News
MLZ is a cooperation between:
 > Technische Universität München
> Technische Universität München > Helmholtz-Zentrum Hereon
> Helmholtz-Zentrum Hereon
 > Forschungszentrum Jülich
> Forschungszentrum Jülich
MLZ is a member of:
 > LENS
> LENS > ERF-AISBL
> ERF-AISBL
MLZ on social media: